Quoted from: https://en.wikipedia.org/wiki/Hierarchical_clustering
Strategies for hierarchical clustering generally fall into two types:
- Agglomerative: This is a "bottom-up" approach: each observation starts in its own cluster, and pairs of clusters are merged as one moves up the hierarchy.
- Divisive: This is a "top-down" approach: all observations start in one cluster, and splits are performed recursively as one moves down the hierarchy.
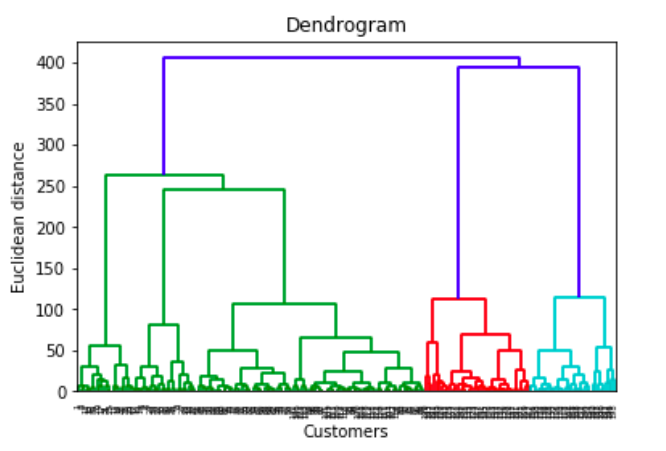
In general, the merges and splits are determined in a greedy manner. The results of hierarchical clustering are usually presented in a dendrogram.
The standard algorithm for hierarchical agglomerative clustering (HAC) has a time complexity of {\displaystyle {\mathcal {O}}(n^{3})} and requires {\displaystyle \Omega (n^{2})}
and requires {\displaystyle \Omega (n^{2})} memory, which makes it too slow for even medium data sets. However, for some special cases, optimal efficient agglomerative methods (of complexity {\displaystyle {\mathcal {O}}(n^{2})}
memory, which makes it too slow for even medium data sets. However, for some special cases, optimal efficient agglomerative methods (of complexity {\displaystyle {\mathcal {O}}(n^{2})} ) are known: SLINK for single-linkage and CLINK for complete-linkage clustering. With a heap, the runtime of the general case can be reduced to {\displaystyle {\mathcal {O}}(n^{2}\log n)}
) are known: SLINK for single-linkage and CLINK for complete-linkage clustering. With a heap, the runtime of the general case can be reduced to {\displaystyle {\mathcal {O}}(n^{2}\log n)} , an improvement on the aforementioned bound of {\displaystyle {\mathcal {O}}(n^{3})}
, an improvement on the aforementioned bound of {\displaystyle {\mathcal {O}}(n^{3})} , at the cost of further increasing the memory requirements. In many cases, the memory overheads of this approach are too large to make it practically usable.
, at the cost of further increasing the memory requirements. In many cases, the memory overheads of this approach are too large to make it practically usable.
Except for the special case of single-linkage, none of the algorithms (except exhaustive search in {\displaystyle {\mathcal {O}}(2^{n})} ) can be guaranteed to find the optimum solution.
) can be guaranteed to find the optimum solution.
Divisive clustering with an exhaustive search is {\displaystyle {\mathcal {O}}(2^{n})} , but it is common to use faster heuristics to choose splits, such as k-means.
, but it is common to use faster heuristics to choose splits, such as k-means.
Metric
The choice of an appropriate metric will influence the shape of the clusters, as some elements may be relatively closer to one another under one metric than another. For example, in two dimensions, under the Manhattan distance metric, the distance between the origin (0,0) and (.5, .5) is the same as the distance between the origin and (0, 1), while under the Euclidean distance metric the latter is strictly greater.
Some commonly used metrics for hierarchical clustering are:
| Names |
Formula |
| Euclidean distance |
{\displaystyle \|a-b\|_{2}={\sqrt {\sum _{i}(a_{i}-b_{i})^{2}}}} |
| Squared Euclidean distance |
{\displaystyle \|a-b\|_{2}^{2}=\sum _{i}(a_{i}-b_{i})^{2}} |
| Manhattan distance |
{\displaystyle \|a-b\|_{1}=\sum _{i}|a_{i}-b_{i}|} |
| Maximum distance |
{\displaystyle \|a-b\|_{\infty }=\max _{i}|a_{i}-b_{i}|} |
| Mahalanobis distance |
{\displaystyle {\sqrt {(a-b)^{\top }S^{-1}(a-b)}}} where S is the Covariance matrix where S is the Covariance matrix |
For text or other non-numeric data, metrics such as the Hamming distance or Levenshtein distance are often used.
A review of cluster analysis in health psychology research found that the most common distance measure in published studies in that research area is the Euclidean distance or the squared Euclidean distance.
Linkage criteria
The linkage criterion determines the distance between sets of observations as a function of the pairwise distances between observations.
Some commonly used linkage criteria between two sets of observations A and B are:
| Names |
Formula |
| Maximum or complete-linkage clustering |
{\displaystyle \max \,\{\,d(a,b):a\in A,\,b\in B\,\}.} |
| Minimum or single-linkage clustering |
{\displaystyle \min \,\{\,d(a,b):a\in A,\,b\in B\,\}.} |
| Unweighted average linkage clustering (or UPGMA) |
{\displaystyle {\frac {1}{|A|\cdot |B|}}\sum _{a\in A}\sum _{b\in B}d(a,b).} |
| Weighted average linkage clustering (or WPGMA) |
{\displaystyle d(i\cup j,k)={\frac {d(i,k)+d(j,k)}{2}}.} |
| Centroid linkage clustering, or UPGMC |
{\displaystyle \|c_{s}-c_{t}\|} where {\displaystyle c_{s}} where {\displaystyle c_{s}} and {\displaystyle c_{t}} and {\displaystyle c_{t}} are the centroids of clusters s and t, respectively. are the centroids of clusters s and t, respectively. |
| Minimum energy clustering |
{\displaystyle {\frac {2}{nm}}\sum _{i,j=1}^{n,m}\|a_{i}-b_{j}\|_{2}-{\frac {1}{n^{2}}}\sum _{i,j=1}^{n}\|a_{i}-a_{j}\|_{2}-{\frac {1}{m^{2}}}\sum _{i,j=1}^{m}\|b_{i}-b_{j}\|_{2}} |
where d is the chosen metric. Other linkage criteria include:
- The sum of all intra-cluster variance.
- The increase in variance for the cluster being merged (Ward's criterion).
- The probability that candidate clusters spawn from the same distribution function (V-linkage).
- The product of in-degree and out-degree on a k-nearest-neighbour graph (graph degree linkage).
- The increment of some cluster descriptor (i.e., a quantity defined for measuring the quality of a cluster) after merging two clusters.